Lab Tools
You can checkout the Wang Bioinformatics Lab code at GitHub.
Mass Spectrometry Query Language¶
The mass spectrometry query language (MassQL) is a domain specific language that specifically aims to express patterns in mass spectrometry data and empower chemists and bioinformaticians to query raw mass spectrometry data. MassQL is designed to be simple, flexible, scalable, and shareable. It aims to make it simple to express a wide range of mass spec data patterns and search across all public mass spectrometry data available, i.e. billions of compounds in hundreds of thousands of samples.
Check it out here.
ModiFinder¶
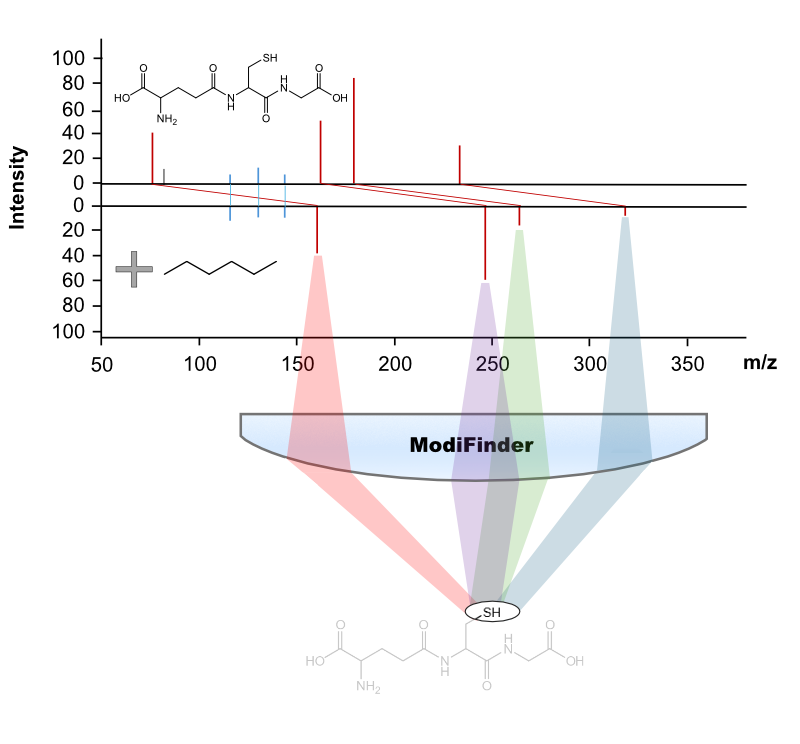
Untargeted tandem mass spectrometry (MS/MS) has emerged as a high-throughput technique for analyzing small molecules in complex samples. A critical objective in this domain is the accurate transformation of MS/MS spectra into chemical structures. While computational methods like MS/MS library searches have facilitated the re-identification of known compounds, and analog library searches alongside molecular networking have extended identification to unknown compounds, there remains a gap in automated methods for pinpointing site-specific structural modifications.

Reza demonstrating his tool to collaborators
To address this challenge, we introduce ModiFinder. This innovative tool leverages the alignment of peaks in MS/MS spectra between structurally related known and unknown small molecules. ModiFinder targets shifted MS/MS fragment peaks during the alignment process, which likely represent substructures of the known molecule containing the modification site. By synthesizing this information, ModiFinder scores the likelihood of each atom in the known molecule being the modification site.
ModiFinder enhances the capabilities of MS/MS analog searching and molecular networking, providing precise localization of modifications and accelerating the discovery of novel compounds. Explore how ModiFinder can transform your research in our detailed demonstration.
Try it out here.
Source Code:
Transitive Alignments¶

Molecular Networks (MNs) have been developed as a computational tool to aid in the organization and visualization of complex chemical space in untargeted mass spectrometry data, thereby supporting comprehensive data analysis and interpretation. MNs group related compounds with potentially similar structures from MS/MS data by calculating all pairwise MS/MS similarities and filtering these connections to produce a MN. Such networks are instrumental in metabolomics for identifying novel metabolites, elucidating metabolic pathways, and even discovering biomarkers for disease. While MS/MS similarity metrics have been explored in the literature, the influence of network topology approaches on MN construction remains unexplored. This manuscript introduces metrics for evaluating MN construction, benchmarks state-of-the-art approaches, and proposes the Transitive Alignments approach to improve MN construction. The Transitive Alignment technique leverages the MN topology to realign MS/MS spectra of related compounds that differ by multiple structural modifications. Combining this Transitive Alignments approach with pseudoclique finding, a method for identifying highly connected groups of nodes in a network, resulted in more complete and higher-quality molecular families. Finally, we also introduce a targeted network construction technique called induced transitive alignments where we demonstrate effectiveness on a real world natural product discovery application. We release this transitive alignment technique as a high-throughput workflow that can be used by the wider research community.
GNPS2¶
GNPS2 is the next generation of the GNPS ecosystem. It is a complete rewrite of the entire GNPS ecosystem to be more scalable, more modular, and more extensible. It is currently in beta testing and if you'd like to contribute or try it out let us know. You can find it here along with its documentation.
GNPS¶
GNPS is an entire analysis ecosystem that is comprises computational workflows, community aggregated knowledge, public repository data, and data visualization tools. Summarized below are just a small set of the tools available in GNPS.
GNPS Spectral Libraries¶
Spectral libraries -- a collection of reference tandem mass spectrometry data from known compounds -- are a principal unit of knowledge within the mass spectrometry community. Using spectral libraries of reference compounds is the most common method to identify known compounds in untargeted experiments. However, spectral libraries traditionally were fragmented across the community, silo'd in individual labs. GNPS spectral libraries created the infrastructure to crowd source and enpower the community to deposit their spectral libraries in a centralized location. This has enable the growth of spectral libraries from a few thousand in 2014 to over 500K in 2022.
Check it out here.
Classical Molecular Networking¶
Molecular Networking is a computational tool that groups up similar MS/MS spectra based upon their fragmentation. The entire pipeline also features multivariate statistics, and spectral library search.
Check it out here.
Feature Based Molecular Networking¶
Feature Based Molecular Networking is a technique that integrates quantitative feature finding tools with molecular networkings. This transforms the qualitative comparisons between conditions with Classical Molecular Networking into a quantitative comparison, where relative abudance can be used to prioritize identification efforts. This is a broad community effort that uniquely is widely compatible across the most popular open source software and many proprietary vendor software.
GNPS Dashboard¶
The GNPS Dashboard is the only fully web based mass spectrometry interactive visualization tool that enables google-docs like collaboration and sharing of results. It is deeply integrated into community resources, including all proteomics, metabolomics, and glycomics public data repositories as well as online analysis systems such as GNPS. GNPS Dashboard drastically lowers the barrier to entry to visualize and interrogate mass spec data from nearly all instruments with a single click, without the need for proprietary software or any local software installation. This makes it perfect for classroom teaching and data transparency for manuscript reviews and post publication inspection.
Check it out here.
ReDU¶
ReDU is a crowd sourcing tool that has uniquely facilitated the annotation of over 50K public metabolomics analyses with sample information using controlled vocabularies. By aggregating all this information, ReDU empowers the community to seemlessly select subsets of public data for reanalysis and metaanalysis.
Check it out here.
MASST¶
The Mass Spectrometry Search Tool (MASST) is the first tool to enable data driven searching of all public metabolomics data at a repository scale. MASST is the analog to BLAST in the sequencing world that enabled the searching of NCBI and SRA with a nucleotide sequence. Here, MASST enables the searching of molecules using their tandem mass spectra across all public mass spectrometry data from all major repositories: MassIVE, Metabolights, and Metabolomics Workbench.
Check it out here.
Miscellaneous Lab Tools¶
Co-author Summarization Tool for NSF Grants: link
Other Tools¶
Checkout other tools in the GNPS2 Platform here.